Since my bachelor thesis I use cloning on a regular basis. Back then to transform yeast, later I needed to clone in order to perform a yeast-two-hybrid screen while now everything has shifted to the expression and modification of proteins.
Cloning of plasmids is easy due to the long known and appreciated restriction enzymes. You flank your coding sequence with unique restriction sites and ligate it into a multiple cloning site of an expression vector. You can attach another promoter, exchange the terminator, and adjust the plasmid to your specific needs. Then you go on using your plasmid to express the protein you designed in a cell the way you want.
But when I want to change parts of a protein, I need to go back to the level of DNA and clone a new plasmid with the according changes in the coding region and run through the whole process of protein expression again. I wished for an enzyme which enables me to exchange a specific part of a protein as I do in plasmids.
Fantastic Sortases and where to find them
At the beginning of my PhD I finally heard of the bacterial enzyme Sortase A. It is a protein that is both restriction enzyme and ligase combined – but in contrast to the classical restriction enzymes, it works on proteins. Like restriction enzymes for DNA the Sortase A recognizes a specific sequence and cuts at a defined site. And like a ligase for DNA it ligates the amino acid of one end back to a matching end.
Why would bacteria need such an enzyme that cuts and ligates other proteins? Well, because bacteria and I have the same problem: We both want a simple to handle tool to exchange parts of proteins and alter their function. Staphylococcus aureus is one example for bacteria using Sortase A, Streptococcus pyogenes another. They are gram-positive bacterial pathogens that use several proteins in their cell wall to prevent detection by a host. For example, they use their immunoglobulin G-binding protein A (Protein A / spa) to bind the Fc domains of the host’s antibodies. Other examples are the fibronectin-binding proteins A and B (FnbA & FnbB), which of course bind the host’s fibronectin. The pathogens thereby cover their surface with host proteins so that antibodies no longer opsonize them. Additionally, these host proteins promote the interaction with the host tissues in order to invade host cells.
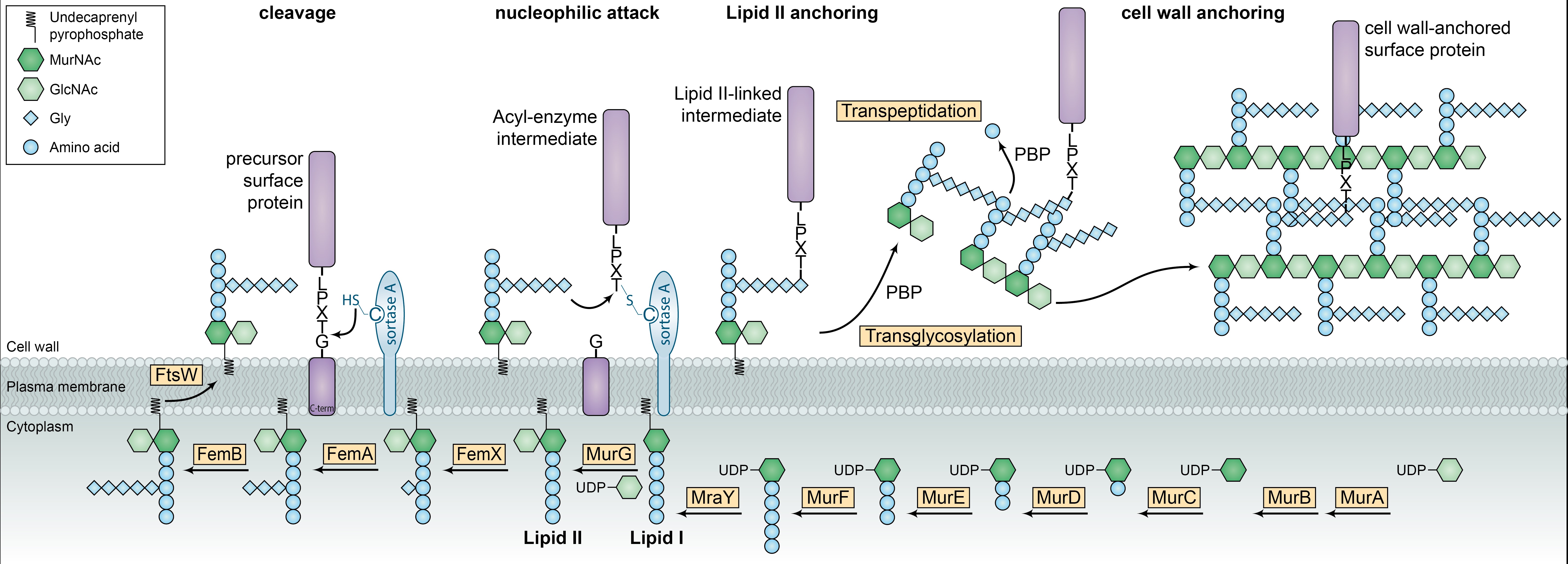
Figure 1) Synthesis of gram-positive bacterial cell wall and anchoring of surface proteins to Lipid II by sortase A. The figure is adapted from Schneewind et al 2011 and Veening et al 2013. (Adapted by permission from Macmillan Publishers Ltd: Nat. Rev. Microbiol., copyright 2011 and Nat. Rev. Microbiol., copyright 2013.)
Bacteria have several options to present proteins on their surface. Proteins with a secretion signal and a transmembrane domain are sorted to the plasma membrane. But in order to reach beyond the peptidoglycan cell wall these proteins need to be big. Proteins can be glycosylated to be linked to sugars of the cell wall by transglycosylation. The glycosylation that can vary in sugars and branching is performed in the Golgi. The glycosylated proteins are then sorted to the plasma membrane to be secreted. The secreted and glycosylated protein is then linked to the sugars of the cell wall. Finally, proteins can be linked to other proteins or peptides in the cell wall by transpeptidation. Again, the protein needs a signal to be secreted. Enzymes like Sortases then link secreted proteins based on a recognition site to peptides of the peptidoglycan cell wall.
As you see, bacteria use different ways to modify proteins to meet their needs. First genetically, which would correspond to the genetic manipulation of coding sequences on plasmids. Then by adding sugars which I am not interested in. And finally, by ligating proteins to specific amino acids. Hurray! Gram-positive bacteria have invented the tool I was looking for.
Fantastic Sortases – Rise of the Extra Super Sortase
Luckily, Hidde Ploegh and colleagues were already developing transpeptidases as biochemical tools. They selected the enzyme Sortase A from Staphylococcus aureus recognizing the peptide sequence LPXTG and the Sortase A from Streptococcus pyogenes recognizing LPXTA and LPXTG. SrtA hydrolyses the peptide bond between Thr and Gly/Ala and ligates it back to any free n-terminal Gly/Ala. In order to develop SrtA to an easy to use tool, they removed the transmembrane domain and mutated five amino acids of the S. aureus SrtA (5M SrtA) to increase the efficiency 50 times. This 5M SrtA was therefore initially called ‘Super Sortase’. And then they mutated two additional amino acids from the Super Sortase and thereby created the 7M SrtA. This enzyme is no longer dependent on magnesium and was initially termed Extra Super Sortase.
Fantastic Sortases and how to use them
In my PhD I use these enzymes on proteins I previously expressed in and purified from mammalian cells. These proteins have an n-terminal hexahistidine tag followed by the SrtA motif LPETG. Using the 5M SrtA I can now remove the n-terminal 6HIS tag and either leave the n-terminus of my protein like it is or attach another protein or small molecule containing the LPETG motif. The method can also be used to attach small molecular dyes or fluorescent proteins to the c-terminus of antibodies. Buy one antibody and attach whatever color you need for your FACS analysis. Plus, the payload is added site specifically and therefore interferes less likely with antibody binding. Moreover, just one payload can be added per LPETG motif resulting in homogenous batches of tagged antibody. It even works on proteins located on the surface of cells. So you could attach a small molecule dye to a receptor and observe its translocation after activation. You can use a Sortase just to cleave proteins where traditionally a TEV protease would be used. The great benefit of using the SrtA is that you keep the option to modify the protein after the cleavage.
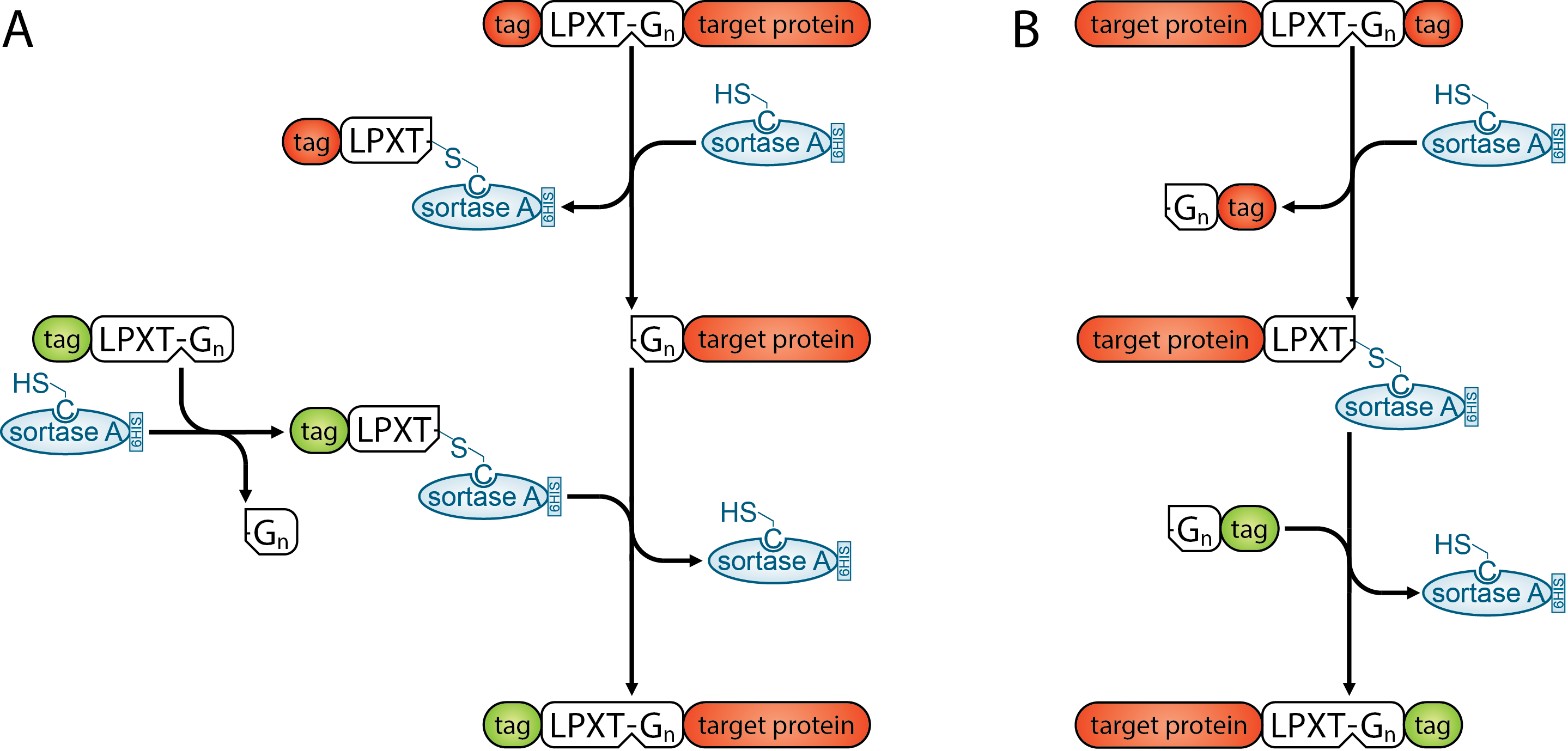
Schematic description of N-terminal (A) and C-terminal (B) transpeptidation by Sortase A. (Figure by S. Görgen.)
So why do I think SrtA is awesome for ligation of peptides to proteins? When used to cleave a protein site specifically, SrtA gives the additional option to modify the protein later. When used to tag a payload to a protein, SrtA adds site specifically and maximally one payload per recognition motif, which offers more control compared to the common chemical tagging methods.
References:
Hendrickx, A.P.A., Budzik, J.M., Oh, S.-Y., and Schneewind, O. (2011). Architects at the bacterial surface – sortases and the assembly of pili with isopeptide bonds. Nat. Rev. Microbiol. 9, 166–176.
Pinho, M.G., Kjos, M., and Veening, J.-W. (2013). How to get (a)round: mechanisms controlling growth and division of coccoid bacteria. Nat. Rev. Microbiol. 11, 601–614.
Simon Görgen
(featured image from colourbox.com)
