
(Rolf Müller/UKB)
It has long been known that recognition of microbial DNA in the cytosol induces an antiviral type I interferon immune response. However, until recently, only long double-stranded DNA had been considered as immune stimulatory. Thus, it has remained unclear whether other structural features than base paired DNA stretches are crucial for activation of cytosolic DNA sensors, and if for example single-stranded reverse transcripts of lentiviruses or endogenous retroelements are also potent interferon stimuli. Now, the group of Dr Martin Schlee and Prof. Gunther Hartmann at the Institute of Clinical Chemistry and Clinical Pharmacology has identified the minimal structural motif required to for single-stranded DNA to be super stimulatory.
Dr Anna Herzner, a postdoc in the Schlee group, together with her co-workers showed that very short dsDNA fragments containing free guanosine ends (G-ended Y-form short DNA (G-YSD)) induce a strong type I interferon response via the cytosolic DNA receptor cGAS. These findings provide important insights into the recognition and a potential viral immune evasion mechanism of HIV-1 as well as the pathology of the autoimmune disease Aicardi–Goutières syndrome (AGS).
Anna, you and your co-authors identified the molecular identity of tiny DNA species causing massive type I interferon induction in a detective masterpiece. Your compelling data was published in Nature Immunology, just a few days ago. Congratulations!
So, can you tell us how the idea was born to search for this kind of immune stimulatory DNA species?
It grew as a project already some time ago. My co-first author Cristina started working on this question. She and Martin were looking for the minimal DNA stimulus that could induce a potent type I interferon response. First, originally the idea was that these DNAs could then be used as an immune stimulatory agent to fight cancer or viruses. Second, the cytosolic DNA sensor was identified very late, in 2013. So there was the hope that this minimal stimulatory DNA motive could serve to identify the cytosolic DNA sensor.
They had started by cutting down the length of dsDNA that still induced a type I interferon response to a minimum of 50 to 60 base pairs (bp). Then, there was a publication from the Paludan lab that showed that single stranded (ss) HIV-derived DNA of only 100 bases was immunostimulatory. In infected cells, HIV first transcribes its RNA genome into ssDNA templates. The very first viral ssDNAs that accumulate in the cytosol are the so-called “strong stop DNAs”. They contain secondary stem loop structures, which were described to be immunostimulatory by the Paludan group. However, they contained short double stranded stretches of only about 30 bp. That these short dsDNA stretches were immunostimulatory was surprising to us, as from our studies these base paired stretches were just too short to stimulate a type I interferon response. But we already knew from Cristina´s work that also single stranded overhangs of dsDNA contribute to cytosolic DNA recognition. So, we were very curious first whether recognition of single stranded parts plays a role in immune stimulation by HIV-1 derived ssDNA and second whether these DNA species can be turned into a highly stimulatory DNA.
How did you discover G-YSD as the highly active, cGAS-stimulatory DNA-species?
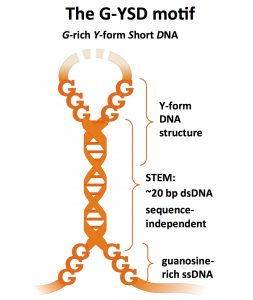
(A. Herzner)
We started by sorting out which stem loop structures within the whole HIV-1 strong-stop DNA was immune stimulatory. We designed individual stem loop structures as synthetic DNAs and tested them on primary human cells. One of these HIV-1 stem loops turned out to be immune stimulatory.
As we had hints that the DNA sequence could play a role, we tested whether the nucleotide sequence influences the immune stimulatory capacity of the HIV-1 derived ssDNA. To our surprise, the stimulatory ssDNA was turned into a non-stimulatory one when depleting guanosines from the single stranded parts.
In order to identify whether mismatches could be responsible for the immune stimulatory capacity of very short dsDNA we depleted mismatches within in the short 20 bp long double stranded part. But this modification actually increased the stimulatory potential of the DNA even further.
When we put all this together we knew that adding guanosines and having a perfect double-strand (free from mismatches in the stem) we gain an optimal DNA stimulus and finally defined a minimal “super-stimulatory” DNA motif as a 20 bp dsDNA flanked by free guanosine ends.
Finally, we showed that the immune stimulatory capacity of our super stimulatory DNAs was dependent on the cytosolic DNA sensor cGAS and that it also activated cGAS in an in vitro assay.
In which context and circumstances do such Y-DNA species occur and in what clinical context do they play a role?
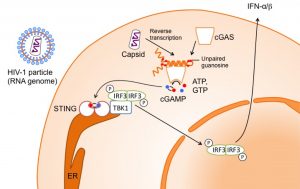
Super-immune stimulatory DNA is involved in the immune response against HIV (image: A. Herzner)
For HIV-1 we proved that mainly ssDNA is present in the cytosol of infected cells. Further, we showed that recognition of such ssDNA species relies on the presence of free guanosines (Gs). Here an amazing fact about HIV is that the HIV-1 ssDNA in fact has a very low G content across all known strains. This likely resembles an evolutionary adaptation of HIV to evade immune recognition. Nevertheless, in some AIDS patients, the so called “elite controllers”, recognition of accumulating HIV-1 ssDNA species is described to protect from onset of the disease.
Interestingly, in the autoimmune disease AGS ssDNA derived from retroelements (endogenous retroviruses which are located everywhere within our genome) accumulates in the cytosol. Based on this, we stimulated cells with retroelement derived ssDNAs containing secondary structures. We saw that an interferon response was induced, and this was also dependent on the presence of single stranded guanosines. This suggests that the recognition of Gs in accumulating ssDNA species is also involved in the interferon mediated pathology present AGS patients.
Altogether, we gained a deeper insight into how retroviruses and retroelements are recognized by our immune system on the molecular level and how HIV-1, for example, might has evolved to evade this host response.

Author: Klara Höning

Pingback: The Cluster goes Christmas – ImmunosensationBlog
Pingback: The Toll Meeting 2015 [toll (German): great, amazing, fantastic] – ImmunosensationBlog